P <- Variable(c(n,n))
ones <- matrix(1, nrow = n, ncol = 1)
obj <- Minimize(lambda_max(P - 1/n))
constr1 <- list(P >= 0, P %*% ones == ones, P == t(P))
constr2 <- list(P[1,3] == 0, P[1,4] == 0)
prob <- Problem(obj, c(constr1, constr2))
result <- psolve(prob)Fastest Mixing Markov Chain
Introduction
This example is derived from the results in Boyd et al. (2004), section 2. Let \(\mathcal{G} = (\mathcal{V}, \mathcal{E})\) be a connected graph with vertices \(\mathcal{V} = \{1,\ldots,n\}\) and edges \(\mathcal{E} \subseteq \mathcal{V} \times \mathcal{V}\). Assume that \((i,i) \in \mathcal{E}\) for all \(i = 1,\ldots,n\), and \((i,j) \in \mathcal{E}\) implies \((j,i) \in \mathcal{E}\). Under these conditions, a discrete-time Markov chain on \(\mathcal{V}\) will have the uniform distribution as one of its equilibrium distributions. We are interested in finding the Markov chain, constructing the transition probability matrix \(P \in {\mathbf R}_+^{n \times n}\), that minimizes its asymptotic convergence rate to the uniform distribution. This is an important problem in Markov chain Monte Carlo (MCMC) simulations, as it directly affects the sampling efficiency of an algorithm.
The asymptotic rate of convergence is determined by the second largest eigenvalue of \(P\), which in our case is \(\mu(P) := \lambda_{\max}(P - \frac{1}{n}{\mathbf 1}{\mathbf 1}^T)\) where \(\lambda_{\max}(A)\) denotes the maximum eigenvalue of \(A\). As \(\mu(P)\) decreases, the mixing rate increases and the Markov chain converges faster to equilibrium. Thus, our optimization problem is
\[ \begin{array}{ll} \underset{P}{\mbox{minimize}} & \lambda_{\max}(P - \frac{1}{n}{\mathbf 1}{\mathbf 1}^T) \\ \mbox{subject to} & P \geq 0, \quad P{\mathbf 1} = {\mathbf 1}, \quad P = P^T \\ & P_{ij} = 0, \quad (i,j) \notin \mathcal{E}. \end{array} \]
The element \(P_{ij}\) of our transition matrix is the probability of moving from state \(i\) to state \(j\). Our assumptions imply that \(P\) is nonnegative, symmetric, and doubly stochastic. The last constraint ensures transitions do not occur between unconnected vertices.
The function \(\lambda_{\max}\) is convex, so this problem is solvable in CVXR. For instance, the code for the Markov chain in Figure 2 below (the triangle plus one edge) is
where we have set \(n = 4\). We could also have specified \(P{\mathbf 1} =
{\mathbf 1}\) with sum_entries(P,1) == 1, which uses the sum_entries atom to represent the row sums.
Example
In order to reproduce some of the examples from Boyd et al. (2004), we create functions to build up the graph, solve the optimization problem and finally display the chain graphically.
## Boyd, Diaconis, and Xiao. SIAM Rev. 46 (2004) pgs. 667-689 at pg. 672
## Form the complementary graph
antiadjacency <- function(g) {
n <- max(as.numeric(names(g))) ## Assumes names are integers starting from 1
setNames(lapply(as.character(1:n), function(v) {
setdiff(1:n, g[[v]])
}), 1:n)
}
## Fastest mixing Markov chain on graph g
FMMC <- function(g, verbose = FALSE) {
a <- antiadjacency(g)
n <- length(names(a))
P <- Variable(c(n, n))
o <- rep(1, n)
objective <- Minimize(cvxr_norm(P - 1.0/n, "2"))
constraints <- list(P %*% o == o, t(P) == P, P >= 0)
## Zero out transition probabilities for non-edges (excluding diagonal)
non_edge_constraints <- do.call(c, lapply(names(a), function(i) {
idx <- as.numeric(i)
js <- setdiff(a[[i]], idx)
lapply(js, function(j) P[idx, j] == 0)
}))
constraints <- c(constraints, non_edge_constraints)
prob <- Problem(objective, constraints)
opt_val <- psolve(prob)
if(verbose)
cat("Status: ", status(prob), ", Optimal Value = ", opt_val, "\n")
list(status = status(prob), value = opt_val, P = value(P))
}
disp_result <- function(states, P, tolerance = 1e-3) {
if(!("markovchain" %in% rownames(installed.packages()))) {
rownames(P) <- states
colnames(P) <- states
print(P)
} else {
P[P < tolerance] <- 0
P <- P/apply(P, 1, sum) ## Normalize so rows sum to exactly 1
mc <- new("markovchain", states = states, transitionMatrix = P)
## Plot igraph directly with padding so self-loops aren't clipped
g <- methods::as(mc, "igraph")
set.seed(42)
lay <- igraph::layout_nicely(g)
lay <- igraph::norm_coords(lay)
pad <- 1.6
par(mar = c(1, 1, 1, 1))
igraph::plot.igraph(g, layout = lay, rescale = FALSE,
xlim = c(-pad, pad), ylim = c(-pad, pad),
edge.arrow.size = 0.5, vertex.size = 20,
edge.label = round(igraph::E(g)$prob, 2),
edge.label.color = "navy", edge.label.cex = 0.9,
vertex.color = "orange", vertex.frame.color = "darkgoldenrod",
vertex.label.color = "navy", vertex.label.font = 2)
}
}Results
Table 1 from Boyd et al. (2004) is reproduced below.
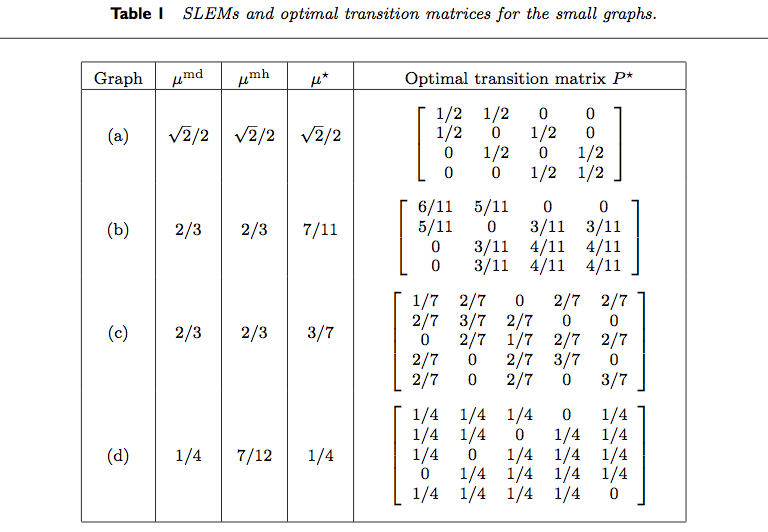
We reproduce the results for various rows of the table.
g <- list("1" = 2, "2" = c(1,3), "3" = c(2,4), "4" = 3)
result1 <- FMMC(g, verbose = TRUE)Status: optimal , Optimal Value = 0.8451543 disp_result(names(g), result1$P)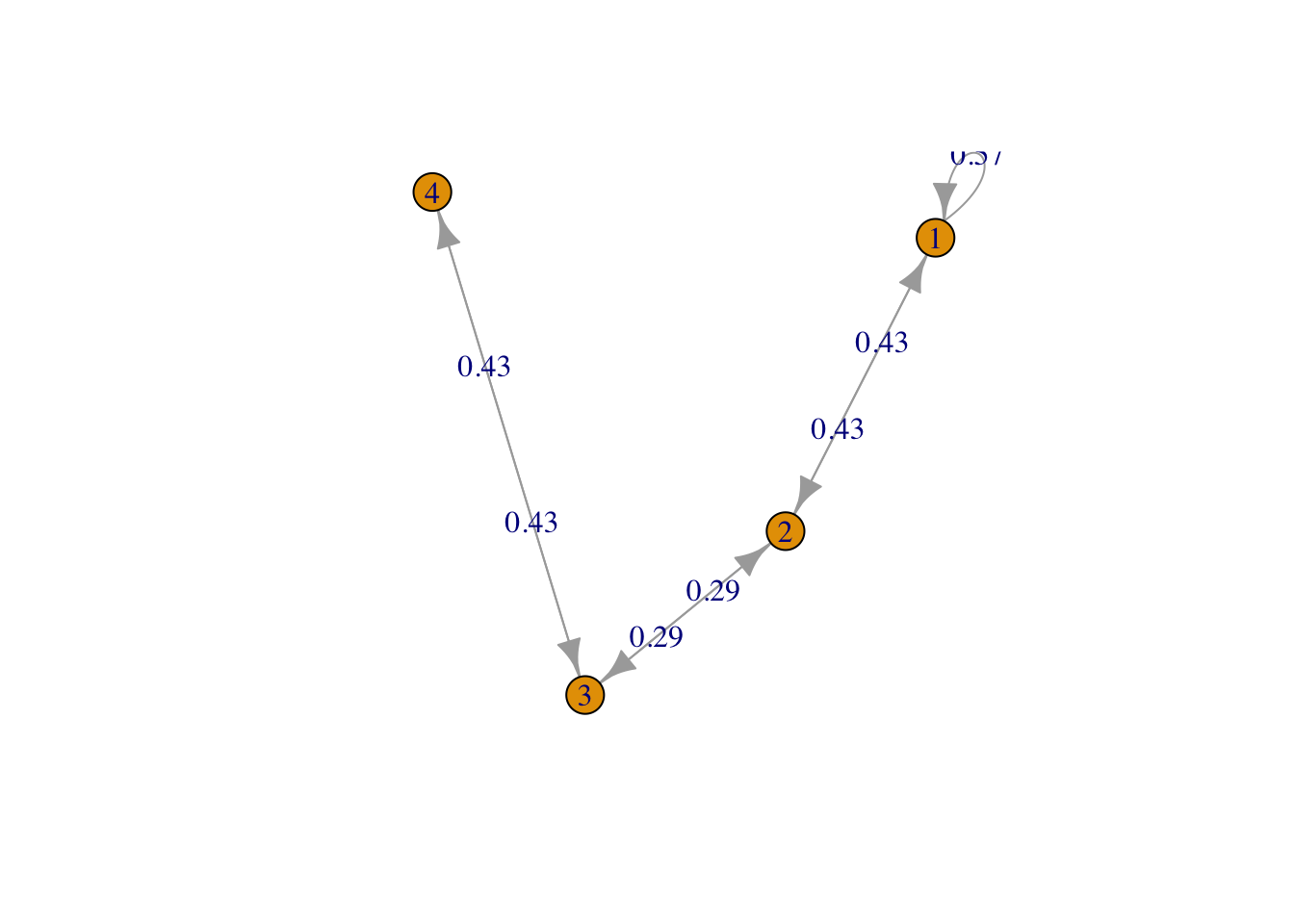
g <- list("1" = 2, "2" = c(1,3,4), "3" = c(2,4), "4" = c(2,3))
result2 <- FMMC(g, verbose = TRUE)Status: optimal , Optimal Value = 0.7071068 disp_result(names(g), result2$P)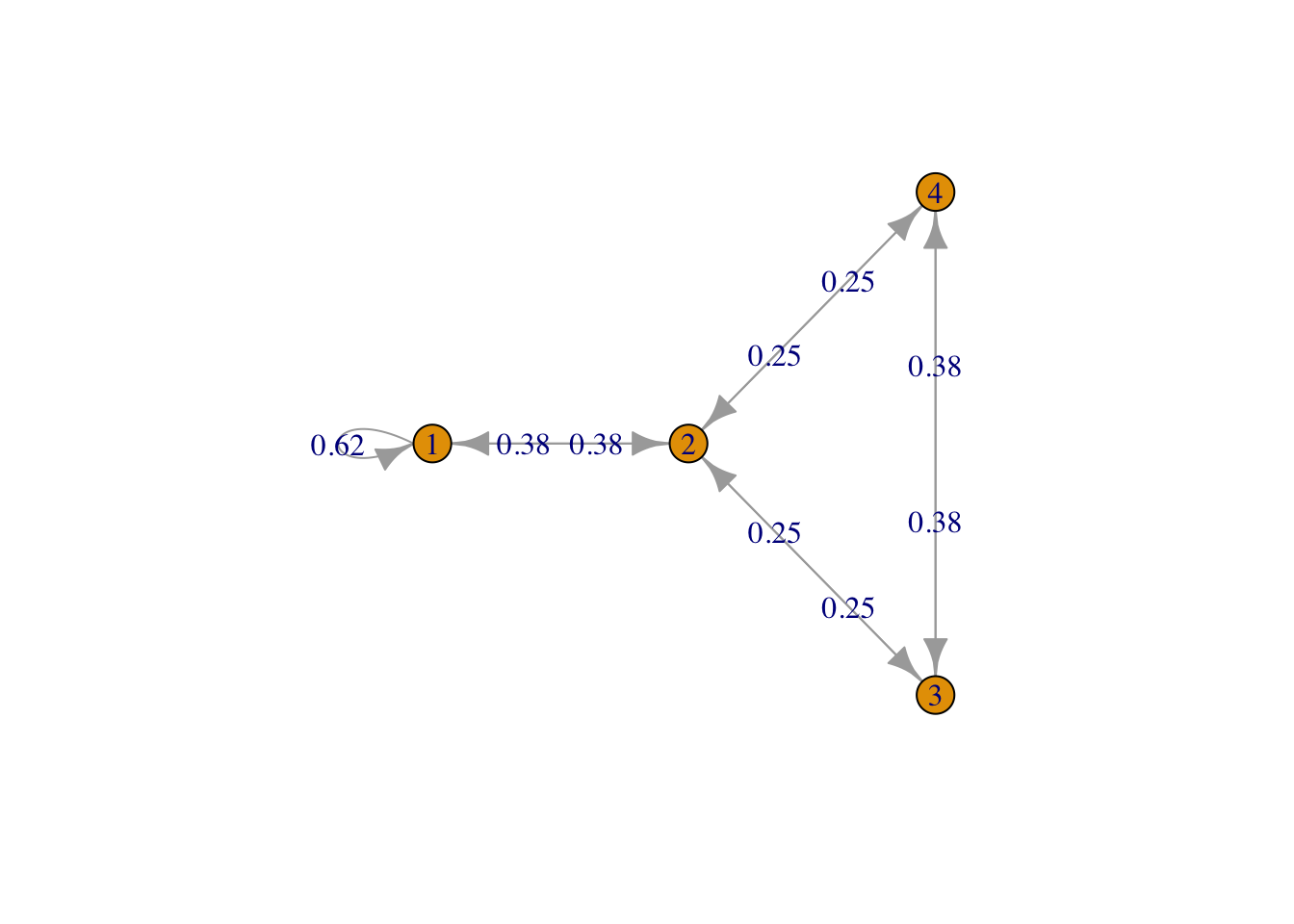
g <- list("1" = c(2,4,5), "2" = c(1,3), "3" = c(2,4,5), "4" = c(1,3), "5" = c(1,3))
result3 <- FMMC(g, verbose = TRUE)Status: optimal , Optimal Value = 0.7559289 disp_result(names(g), result3$P)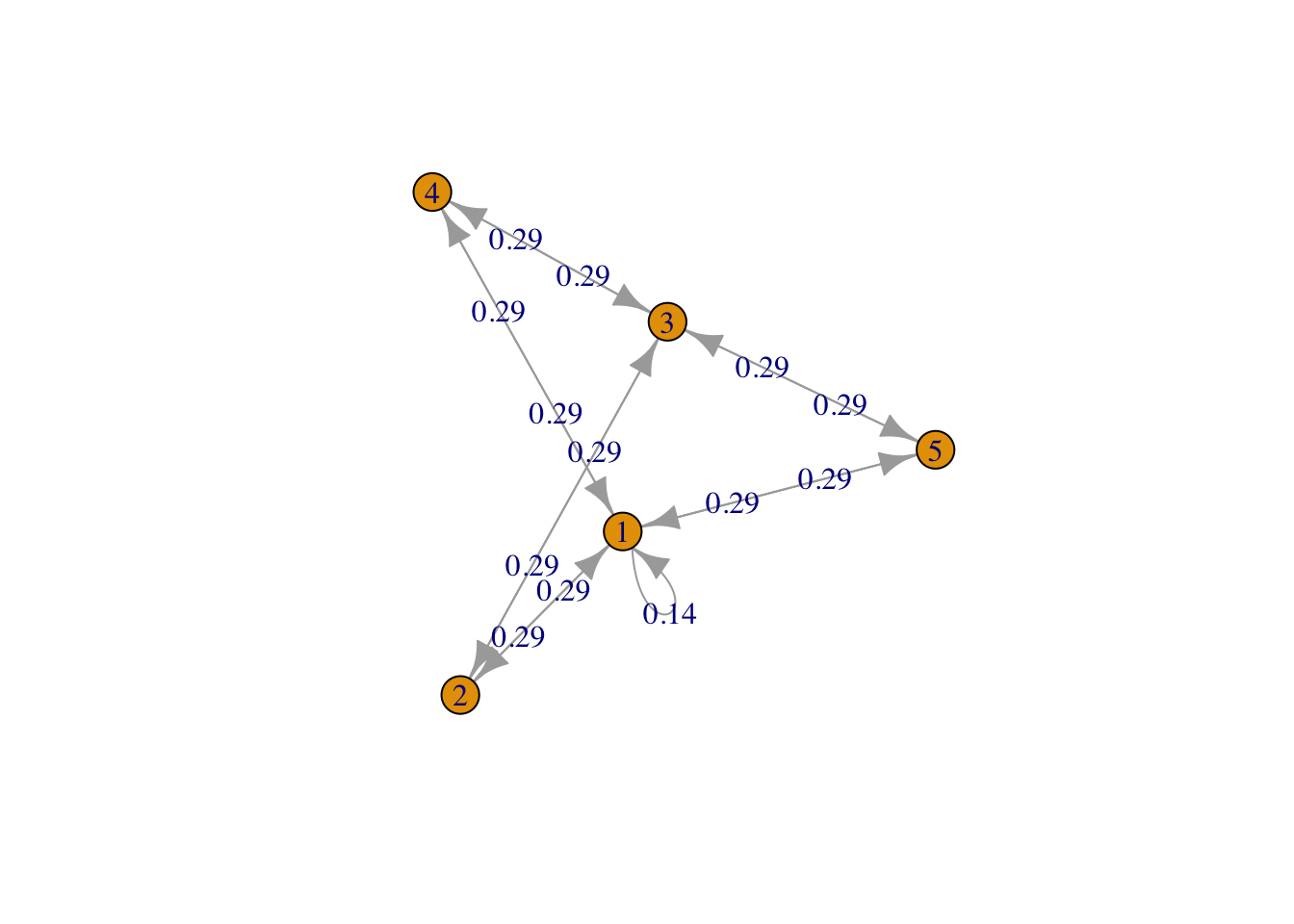
g <- list("1" = c(2,3,5), "2" = c(1,4,5), "3" = c(1,4,5), "4" = c(2,3,5), "5" = c(1,2,3,4,5))
result4 <- FMMC(g, verbose = TRUE)Status: optimal , Optimal Value = 0.4588315 disp_result(names(g), result4$P)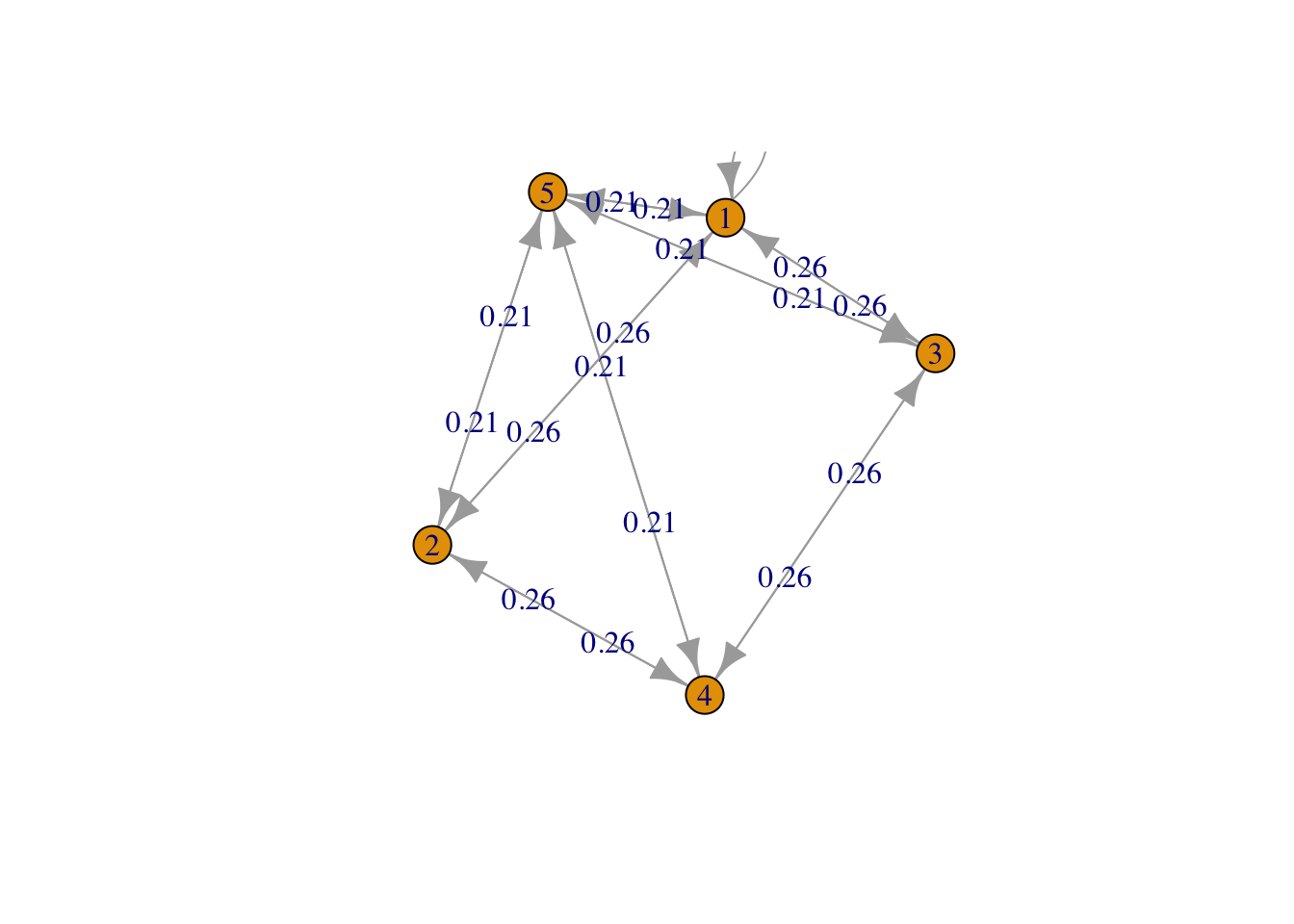
Extensions
It is easy to extend this example to other Markov chains. To change the number of vertices, we would simply modify n, and to add or remove edges, we need only alter the constraints in constr2. For instance, the bipartite chain in Figure 3 is produced by setting \(n = 5\) and
constr2 <- list(P[1,3] == 0, P[2,4] == 0, P[2,5] == 0, P[4,5] == 0)Session Info
R version 4.5.3 (2026-03-11)
Platform: aarch64-apple-darwin20
Running under: macOS Tahoe 26.3.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: America/Los_Angeles
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] markovchain_0.10.3 Matrix_1.7-5 CVXR_1.8.2
loaded via a namespace (and not attached):
[1] piqp_0.6.2 expm_1.0-0 jsonlite_2.0.0
[4] compiler_4.5.3 highs_1.12.0-3 Rcpp_1.1.1
[7] slam_0.1-55 parallel_4.5.3 cccp_0.3-3
[10] yaml_2.3.12 fastmap_1.2.0 clarabel_0.11.2
[13] lattice_0.22-9 igraph_2.2.2 scip_1.10.0-3
[16] knitr_1.51 htmlwidgets_1.6.4 backports_1.5.1
[19] Rcplex_0.3-8 checkmate_2.3.4 gurobi_13.0-1
[22] osqp_1.0.0 rlang_1.1.7 xfun_0.57
[25] S7_0.2.1 RcppParallel_5.1.11-2 otel_0.2.0
[28] cli_3.6.5 magrittr_2.0.4 Rglpk_0.6-5.1
[31] digest_0.6.39 grid_4.5.3 xpress_9.8.1
[34] gmp_0.7-5.1 lifecycle_1.0.5 ECOSolveR_0.6.1
[37] scs_3.2.7 evaluate_1.0.5 codetools_0.2-20
[40] Rmosek_11.1.1 stats4_4.5.3 rmarkdown_2.31
[43] tools_4.5.3 pkgconfig_2.0.3 htmltools_0.5.9